 
Upper triangular half is used) and must be square with dimension equal to If adjacency is a matrix, it is assumed to be symmetric (only the Of each group X (or the last two dimensions if X is 2D). If None, a regular lattice adjacency is assumed, connectingĮach location to its neighbor(s) along the last dimension No adjacency (each location is treated as independent and unconnected). Spatial vertices, frequency bins, time points, etc. adjacency | None | Falseĭefines adjacency between locations in the data, where “locations” can be stat_fun callable() | Noneįunction called to calculate the test statistic. If tail is 0, the statistic is thresholded on both sides of If tail is -1, the statistic is thresholded below threshold. If tail is 1, the statistic is thresholded above threshold. See Notes for an example on how to compute a threshold based onĪ particular p-value for one-tailed or two-tailed tests. If threshold is aĭict (with keys 'start' and 'step') then threshold-freeĬluster enhancement (TFCE) will be used (see the Observations (only valid when using an F-statistic). If None, an F-threshold will be chosenĪutomatically that corresponds to a p-value of 0.05 for the given number of If numeric, vertices with data values more extreme than threshold willīe used to form clusters. (note: this is not an alpha level / “p-value”). The so-called “cluster forming threshold” in the form of a test statistic (observations, frequencies, channels/vertices)). (e.g., spectral data should be provided as Last dimension of each element of X should correspond to theĭimension represented in the adjacency parameter X has 2 groups with respectively 20 and 17 observations in each,Īnd each data point is of shape (50, 4). Number of observations from that group remaining dimensions comprise Parameters : X list of array, shape (n_observations, p) Generated with random partitions of the data. 
Time series, 3D arrays for time-frequency power values). Should contain the data for one group of observations (e.g., 2D arrays for Permutations and cluster-level correction. 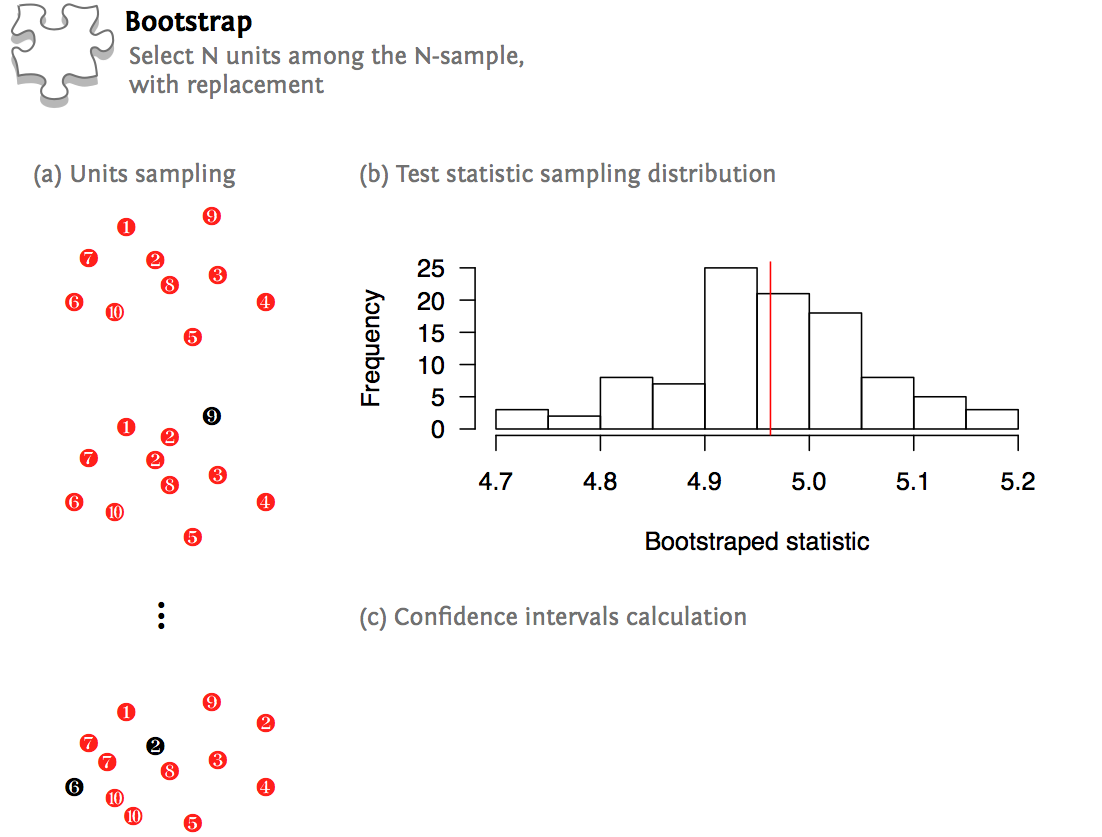
permutation_cluster_test ( X, threshold = None, n_permutations = 1024, tail = 0, stat_fun = None, adjacency = None, n_jobs = None, seed = None, max_step = 1, exclude = None, step_down_p = 0, t_power = 1, out_type = 'indices', check_disjoint = False, buffer_size = 1000, verbose = None ) #Ĭluster-level statistical permutation test.Ĭalculate some statistics corrected for multiple comparisons using 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
